Search
About Anopheles gambiae
Range
Anopheles gambiae senso stricto is the primary mosquito vector responsible for the transmission of malaria in most of sub-Saharan Africa. It is a member of a species complex that includes at least seven morphologically indistinguishable species in the Series Pyretophorus in the Anopheles subgenus Cellia. Anopheles gambiae feeds preferentially on humans and is one of the most efficient malaria vectors known. Anopheles gambiae senso stricto is now known to consist of two genetically distinct forms or incipient species, known formally as the A. gambiae M and A. gambiae S forms. Colonies of these two forms have also been sequenced, assembled and provided here on VectorBase as the A. gambiae Mali-NIH (M) and A. gambiae Pimperena (S) genomes.
Habitats
An. gambiae larvae are generally considered to typically inhabit sunlit, shallow, temporary bodies of fresh water such as ground depressions, puddles, pools and hoof prints . This characteristic may allow predator avoidance as the larvae are able to develop very quickly (~six days from egg to adult under optimal conditions), possibly in response to the ephemeral nature of such larval habitats. An. gambiae larval habitats are therefore often described as containing no (or very sparse) vegetation due to their temporary nature but the great diversity of habitats utilised by An. gambiae includes vegetated (e.g. rice fields) sites. An. gambiae larvae have been reported from habitats containing floating and submerged algae, emergent grass, rice, or 'short plants' and from sites devoid of any vegetation, The variability of larval habitats can be related to the known forms of An. gambiae (e.g. M and S, or Forest, Bamako, Savanna, Mopti and Bissau). For example, the Mopti and M forms are associated with semi-permanent, often man-made, larval habitats such as rice fields or flooded areas, whereas the Savanna/Bamako and S forms are seen more commonly in temporary, rain-dependent sites such as ground puddles.
Resting and feeding preferences
An. gambiae is highly anthropophilic, however, there are indications that An. gambiae can be less discriminant and more opportunistic in its host selection and that host choice is highly influenced by location, host availability and the genetic make-up of the mosquito population. Females of An. gambiae typically feed late at night and are often described as both endophagic and endophilic. Yet there is evidence that indoor and outoor biting are common and both indoor and outdoor resting behaviour appear to be regularly reported. For example, in southern Sierra Leone strong exophily has been demonstrated, linked to the Forest form. Conversely, endophilic behaviour has been linked to Savannah forms. As with host preference, this species appears to exhibit phenotypic plasticity and opportunism in resting locations.
Vectorial capacity
An. gambiae is considered to be one of the most efficient vectors of malaria in the world.
This text was modified from Sinka ME et al. (2010) The dominant Anopheles vectors of human malaria in Africa, Europe and the Middle East: occurrence data, distribution maps and bionomic précis Parasites & Vectors 3:117.
PEST strain
The Anopheles gambiae PEST strain was chosen for genome sequencing because it had both a fixed, standard chromosomal arrangement and a sex-linked pink eye mutation that could readily be used as an indicator of cross-colony contamination (Holt et al 2002: PMID 12364791). The pink eye mutation originated in a colony called A. gambiae LPE established in 1951 at the London School of Hygiene and Tropical Medicine from mosquitoes collected in Lagos, Nigeria. In 1986, this mutation was introduced into a colony of A. gambiae from Asembo Bay western Kenya by crossing males of the LPE strain with female offspring of wild caught Kenyan A. gambiae (the Savanna form), selecting males from the F2 of this cross and then crossing them again with additional female offspring of wild caught Kenyan A. gambiae. From the F2 offspring of this second outcross to Kenyan mosquitoes, a strain was selected that was fixed for pink eye. This outcrossing scheme was repeated one more time in 1987 producing a pink eye strain with a genetic composition largely constituted of the western Kenya Savanna cytogenetic form. In each of these crosses, several hundred female offspring of at least 20 wild caught mosquitoes were used in the cross. This strain, designated A. gambiae PE (Pink Eye), was polymorphic for the inversions 2La (32%) and 2Rbc (19%). The 2Rbc inversion is characteristic of the Mopti chromosomal form, indicating that the original LPE strain from Nigeria was the Mopti form, which is the M molecular form. This inversion was apparently balanced by the uninverted form, because no 2Rbc/bc individuals were detected in the colony. Mukabayire and Besansky selected from this PE strain a set of 9 families whose female parent and at least 20 female offspring were fixed for the standard chromosome karyotype. The progeny of these nine families were pooled to form the A. gambiae PEST strain (Pink Eye STandard). This strain clearly had some Mopti-derived DNA, as the standard karyotype is shared by Mopti and Savanna and the original PE strain did have the 2Rbc inversion rather than 2Rb that is typical of Savanna. Clones from two different PEST strain BAC (Bacterial Artificial Chromosome) libraries had already been end sequenced and physically mapped. When tested, this colony was fully susceptible to P. falciparum from western Kenya. DNA preparation and library construction methods were conducted following standard protocols, and the sequencing method was whole-genome Sanger sequencing. Subsequent to sequencing, the PEST strain was found to be polymorphic for molecular markers diagnostic of the A. gambiae M and S molecular forms. The last known isolate of the A. gambiae PEST colony was lost in about 2005.
Source: VectorBase
Picture credit (public domain): James Gathany (CDC) 1994
Taxonomy ID 7165
Data source VectorBase
Comparative genomics
What can I find? Homologues, gene trees, and whole genome alignments across multiple species.
 More about comparative analyses
More about comparative analyses
 Phylogenetic overview of gene families
Phylogenetic overview of gene families
 Download alignments (EMF)
Download alignments (EMF)
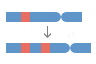
Variation
What can I find? Short sequence variants and longer structural variants.
 More about variation in Ensembl Metazoa
More about variation in Ensembl Metazoa




 Display your data in Ensembl Metazoa
Display your data in Ensembl Metazoa
 Update your old Ensembl IDs
Update your old Ensembl IDs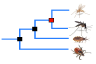
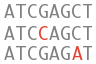

![Follow us on Twitter! [twitter logo]](/i/twitter.png)
