Search
Bemisia tabaci Uganda 1
Whiteflies of the Bemisia tabaci (Gennadius) (Hemiptera: Aleyrodidae) species complex are phloem-feeding insects and plant-virus vectors, some of which are widely regarded to be amongst the world’s worst agricultural pests. Outbreaks of B. tabaci cause significant crop losses and contribute to global food insecurity.
The whitefly Bemisia tabaci Uganda 1, also referred to as Bemisia Uganda1, B. tabaci Uganda, B. tabaci Uganda sweet potato and B. tabaci Uganda 8, has been reported from a range of vegetable and weed hosts in Uganda [1-3]. It has not been reported from cassava or in association with cassava disease epidemics [1-4]. The plant hosts this whitefly has been reported from include: Ipomoea batatas (sweetpotato), Solanum lycopersicum (tomato), Commelina benghalensis (wandering Jew), Leonotis nepetifolia (lion's ear), Pavonia urens (muwugula), Cleome gynandra (joobyo), Vernonia amygdalina (mululuza) and Euphorbia heterophylla (Mexican fire plant) [1-3].
Recent phylogenetic analysis, based on partial mitochondrial COI gene sequences, showed that B. tabaci Uganda 1 grouped outside of the B. tabaci cryptic species complex, basal to the monophyletic group of B. tabaci species [1]. Adults of this species, however, are morphologically indistinguishable from other B. tabaci species [1]. As well as being phylogenetically distinct from the sub-Saharan African cassava B. tabaci species, it is likely that B. tabaci Uganda 1 transmit different groups of plant-viruses that constrain sweetpotato production Uganda [5,6].
The genome described here was generated from a Ugandan population of B. tabaci Uganda 1, that was inbred in the laboratory to reduce heterozygosity.
The Bemisia tabaci cryptic species complex
Members of the B. tabaci species complex cause plant damage by feeding on plant-phloem sap, inducing phytotoxic disorders, depositing honeydew on which sooty moulds develop and by vectoring > 300 plant-virus species in the genera Begomovirus, Carlavirus, Crinivirus, Ipomovirus, Polerovirus and Torradovirus [7,8]. Diseases caused by these viruses often spread rapidly with devastating yield losses of up to 100% [9].
Bemisia tabaci sensu lato currently represents a relatively large group (>44) of mostly unresolved cryptic species, as inferred from phylogenetic species delimitation studies [10,11]. These morphologically indistinguishable species differ from one another not only in their genetic relatedness, but also in various biological traits such as plant host-range breadth, fecundity, insecticide resistance, and plant-virus transmission efficiencies.
Bemisia tabaci sensu lato are distributed globally, from tropical to temperate climatic zones and across all continents (except Antarctica) [11]. Most cryptic species in this complex, as currently understood, are geographically restricted, but two of them are highly invasive globally i.e., B. tabaci Middle East-Asia Minor 1 (MEAM1, also referred to as biotype B and Bemisia argentifolii) and B. tabaci Mediterranean (MED, also referred to as biotype Q) [11]. Bemisia tabaci sensu lato live predominantly on herbaceous plant hosts and have been recorded from an exceedingly broad range of host plants (>500 species) [12]. The documented host-plant range of most cryptic species within the complex remains largely incomplete.
Picture credit: Sharon van Brunschot.
Taxonomy ID 7038
Data source Ensembl Metazoa
Comparative genomics
What can I find? Homologues, gene trees, and whole genome alignments across multiple species.
 More about comparative analyses
More about comparative analyses
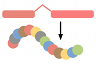
 Phylogenetic overview of gene families
Phylogenetic overview of gene families
 Download alignments (EMF)
Download alignments (EMF)
Variation
This species currently has no variation database. However you can process your own variants using the Variant Effect Predictor:




 Display your data in Ensembl Metazoa
Display your data in Ensembl Metazoa
 Update your old Ensembl IDs
Update your old Ensembl IDs

![Follow us on Twitter! [twitter logo]](/i/twitter.png)
